Results with checkpoint inhibitors nivolumab (PD-1, Opdivo), pembrolizumab (PD-1, Keytruda), and atezolizumab (PD-L1, Tecentriq) are impressive. Some patients have experienced incredible and prolonged responses. These drugs are truly modern medical breakthroughs.
Even patients with large tumor burdens and many mutations have experienced excellent results. The same is true for patients with high amounts of circulating tumor (ct) DNA as determined by “liquid biopsy.” In a study of mutation burden and response, patients with very high tumor burden had the highest response rate to immunotherapy:
This makes sense because patients with many mutations generate neoantigens to which the immune system can mount a response. It is likely that the immune response to at least one of the neoantigens has not been abrogated. Therefore, checkpoint inhibition, which blocks the immune system from no longer attacking these antigens, would be beneficial. Indeed, patients with mutagen-induced tumors, for example, melanoma (UV exposure) and non-small cell lung cancer (cigarette smoking) experience better responses than those with non-mutagen induced tumors.
However, most patients who receive treatment with checkpoint inhibitors, do not experience spectacular responses. What is more troubling is that some patients actually progress rapidly once checkpoint inhibition therapy is initiated. These patients are called “hyperprogressors,” and it is essential to identify hyperprogressors before initiating treatment.
What factors have been associated with hyperprogression?
Approximately 9% of patients across many tumor types are hyperprogressors. Baseline tumor burden and tumor type were not correlated with hyperprogression, however, older age patients were more likely to hyperprogress on anti-PD-1/PD-L1 therapy than younger patients.
The pace of progression in patients with amplification (increased copy number) of the MDM2 gene, which encodes p53, has been shown to increase five to forty-fold compared with those in whom MDM2 amplification is not present. The MDM2 gene encodes a protein that suppresses p53 expression. MDM2 binds to the p53 transactivation domain and repress its transcriptional activity. p53 is the guardian of the genome; it is responsible for pausing replication so that DNA repair can take place, and if repair is not possible, p53 invokes apoptosis. MDM2 amplification deprives cancer cells of this safeguard.
Independent of the affects mediated through p53 suppression, MDM2 has several critical functions that promote invasiveness and genomic instability, and suppress apoptosis.
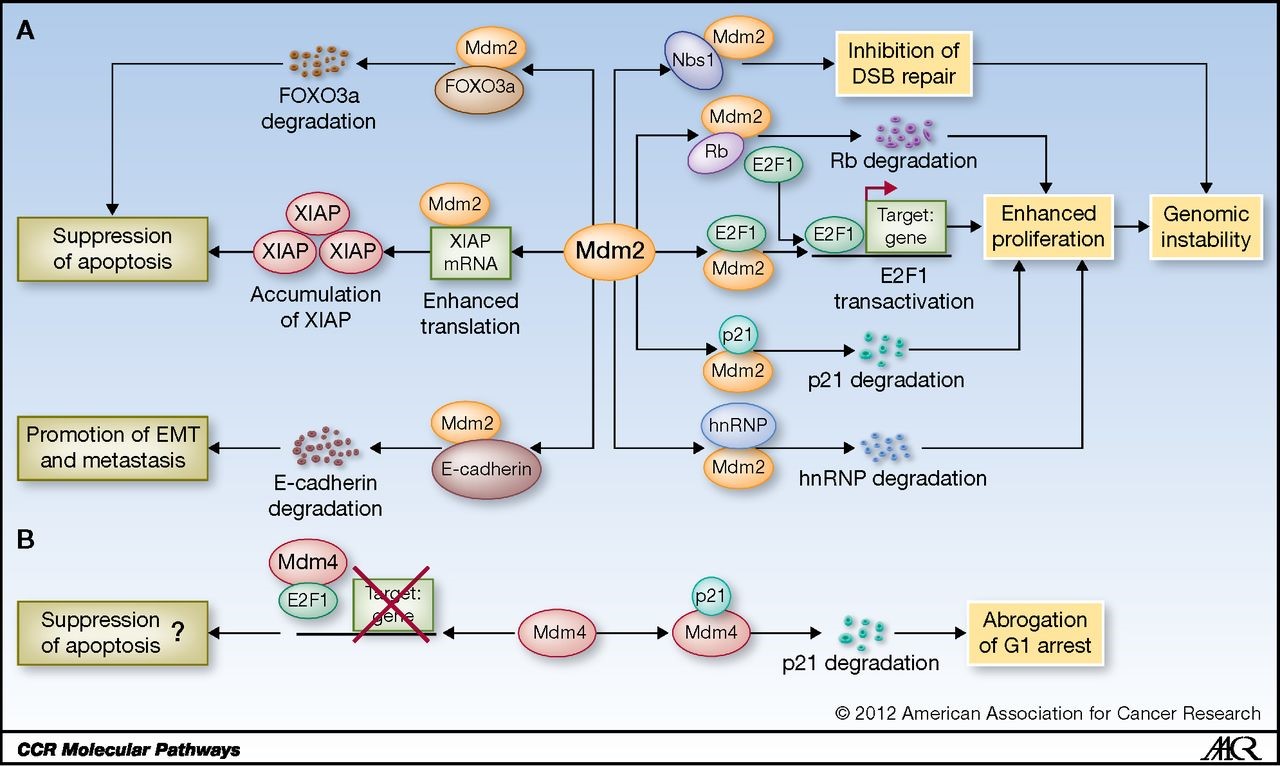
Figure 1. Schematic representation of p53-independent functions of Mdm2 and Mdm4. A, Mdm2 interacts with multiple factors to affect genomic instability, apoptosis, and metastasis. hnRNP, hnRNP K protein. B, Mdm4 interacts with p21 and E2F1 to regulate cell-cycle arrest and apoptosis. http://clincancerres.aacrjournals.org/content/19/1/34
Prior to the availability of checkpoint inhibitors, MDM2 amplification has been associated with poor progression in patients with lung cancer:
Molecular data were correlated with clinicopathological factors and evaluated for their prognostic value. The study group included 116 NSCLC patients who underwent pulmonary resection between 1996 and 1999. MDM2 amplification was found in 24 patients (21%). There was no relationship between MDM2 amplification and clinicopathological factors, such as sex, age and stage of disease, pT, pN, histology and tumor differentiation. Median disease-free survival (DFS) in patients with and without MDM2 amplification was 3 and 31 months, and 5-year DFS 24 and 33%, respectively (log-rank, P=0.02). Likewise, median overall survival (OS) in patients with and without MDM2 amplification was 9 and 33 months, respectively, and 5-year OS 24 and 39%, respectively (log-rank, P=0.01). The strong prognostic relevance of MDM2 amplification for both DFS and OS was confirmed in multivariate analysis (P<0.01 for both comparisons).
In colon cancer, MDM2 amplification, but NOT the single nucleotide polymorphism (thymine to guanine) at codon 72 (SNP309), was associated with advanced tumor stage:
MDM2 was amplified in 9% of the 284 colorectal cancers analyzed and a significantly higher proportion of tumors with high MDM2 gene amplification retained a wild-type p53 gene (P = 0.058). MDM2 gene amplification was significantly correlated to advanced tumor stage.

