Hello all and welcome to this corner of the Interprofessional Health Sciences Library and Information Commons website, an area devoted to bringing you what we hope you find interesting and exciting stories of innovation in biomedical science. Thank you for joining us!
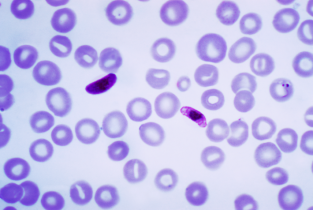
This first installment focuses on malaria – a worldwide health problem that, according to the Centers for Disease Control and Prevention, killed some 445,000 people in 2016. Malaria is caused by parasites, carried by mosquitos which introduce the microorganisms into humans’ blood. Infection results in flu-like fever/chills, headache, and respiratory problems. Left untreated – progression to more serious and life-threatening complications is rapid. The parasite most responsible for this scourge is the protozoan Plasmodium falciparum. Pharmacological interventions exist – but have been thwarted by the unicellular organisms’ ability to acquire resistance. For example, chloroquine and related compounds were employed for decades to disable the parasite’s lysosome-like digestive vacuole. With the organism unable to completely metabolize the host red blood cells’ hemoglobin, toxic metabolites amass and the parasites die. The more recent drug of choice is artemisinin, a natural product derived from the plant Artemisia annua. Compounds containing artemisinin, or related semi-synthetic derivatives, reduce the number of parasites in the bloodstream through what is thought to involve formation of highly reactive free radical species. Unfortunately, identification of other safe drugs with efficacy in killing the microorganism has not been successful.
Which brings us to the new approach – pioneered by a team of investigators of the University of South Florida and Wellcome Trust Sanger Institute and described in a research article entitled “Uncovering the essential genes of the human malarial parasite Plasmodium falciparum by saturation mutagenesis”[1]. Here, a technique called piggyback transposition mutagenesis was used to insert crippling DNA stretches into genes across the organism’s genome. Non-essential genes were so-identified by their ability to withstand the random insertions. Essential genes, in contrast, resulted in the parasite’s death. Illumina-based deep sequencing allowed for identification of the insertion sites – and the analysis was on!
Of the 5399 genes that exist in the P. falciparum genome, 2680 were found to be essential for the parasite’s disease propagation processes. Powerful bioinformatic analyses revealed that essential gene clusters exist encoding proteins involved in, among other functions – lipid metabolic pathways, proteasomal degradation, cell cycle control, and RNA stability. Naturally, these genes and their associated proteins/pathways now are the molecular targets for future antimalarial drug development. It is certain that sophisticated drug screens are currently being employed and we can only hope that safe and efficacious compounds are found in a timely manner. With the dearth of currently available antimalarials and the disease’s continued health toll, advances cannot come too soon.
SRT – June 2018
Reference:
[1] M. Zhang et al., Science 360, eaap7847 (2018). DOI: 10:1126/science.aap7847 (Link to Text)
